Rapport de calcul
Carte Horaire
Compte rendu de calcul Bernese GPS Software
 Informations sur le processus
Informations sur le processus Sessions intégrées au calcul
Sessions intégrées au calcul Orbites et coordonnées du pôle
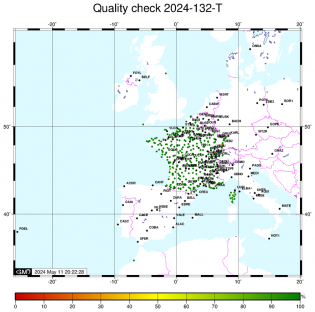
Orbites et coordonnées du pôle Controle qualité
Controle qualité Calcul sur le code
Calcul sur le code Fixation des ambiguités
Fixation des ambiguités Fixation des ambiguités Core
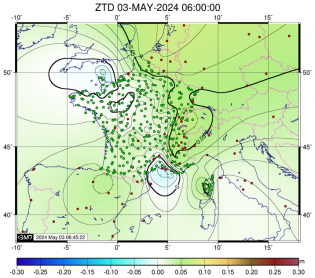
Fixation des ambiguités Core Estimation des ZTD finaux (coordonnées fixées) - ADDNEQ2
Estimation des ZTD finaux (coordonnées fixées) - ADDNEQ2



Sessions intégrées au calcul
| Année | DOY | Session |
|---|---|---|
| 2026 | 136 | r |
| 2026 | 136 | s |
| 2026 | 136 | t |
| 2026 | 136 | u |
| 2026 | 136 | v |
| 2026 | 136 | w |




Orbites et coordonnées du pôle
| Orbites | Coordonnées du pôle |
|---|---|
| IGS0OPSULT_20261351800_02D_15M_ORB.SP3.gz | IGS0OPSULT_20261351800_02D_01D_ERP.ERP.gz |




Controle qualité
| Station | Dernier fichier | Ratio des fichiers précédents | Ratio qc_gps | Ratio qc_glo | qc - header GPS | qc - header GLO | IOD slips <10° | IOD slips >10° | Station utilisée dans le calcul |
|---|---|---|---|---|---|---|---|---|---|
| AJAC | - | 0.0% | - | - | - | - | - | - | - |
| ALPE | - | 100.0% | - | - | - | - | - | - | - |
| ANNY | - | 0.0% | - | - | - | - | - | - | - |
| ARUF | - | 100.0% | - | - | - | - | - | - | - |
| BANN | - | 100.0% | - | - | - | - | - | - | - |
| BRE2 | - | 0.0% | - | - | - | - | - | - | - |
| BRET | - | 0.0% | - | - | - | - | - | - | - |
| BVOI | - | 100.0% | - | - | - | - | - | - | - |
| BVS2 | - | 100.0% | - | - | - | - | - | - | - |
| CABW | - | 0.0% | - | - | - | - | - | - | - |
| CAMO | - | 0.0% | - | - | - | - | - | - | - |
| CAUS | - | 0.0% | - | - | - | - | - | - | - |
| CH2T | - | 100.0% | - | - | - | - | - | - | - |
| CHBL | - | 100.0% | - | - | - | - | - | - | - |
| CHE2 | - | 100.0% | - | - | - | - | - | - | - |
| CHOL | - | 100.0% | - | - | - | - | - | - | - |
| CHPO | - | 100.0% | - | - | - | - | - | - | - |
| CHTL | - | 0.0% | - | - | - | - | - | - | - |
| COCR | - | 100.0% | - | - | - | - | - | - | - |
| CONN | - | 100.0% | - | - | - | - | - | - | - |
| CORZ | - | 100.0% | - | - | - | - | - | - | - |
| COUT | - | 0.0% | - | - | - | - | - | - | - |
| CRAU | - | 100.0% | - | - | - | - | - | - | - |
| CRTS | - | 100.0% | - | - | - | - | - | - | - |
| CSTO | - | 100.0% | - | - | - | - | - | - | - |
| CULA | - | 100.0% | - | - | - | - | - | - | - |
| CYLM | - | 100.0% | - | - | - | - | - | - | - |
| DBMH | - | 100.0% | - | - | - | - | - | - | - |
| DENT | - | 0.0% | - | - | - | - | - | - | - |
| DEQU | - | 100.0% | - | - | - | - | - | - | - |
| DHUI | - | 100.0% | - | - | - | - | - | - | - |
| DOCX | - | 100.0% | - | - | - | - | - | - | - |
| DOMP | - | 100.0% | - | - | - | - | - | - | - |
| DRUL | - | 100.0% | - | - | - | - | - | - | - |
| EBN2 | - | 100.0% | - | - | - | - | - | - | - |
| EQH2 | - | 100.0% | - | - | - | - | - | - | - |
| FEUR | - | 100.0% | - | - | - | - | - | - | - |
| FJC2 | - | 0.0% | - | - | - | - | - | - | - |
| GAJN | - | 100.0% | - | - | - | - | - | - | - |
| GARD | - | 100.0% | - | - | - | - | - | - | - |
| GIE1 | - | 100.0% | - | - | - | - | - | - | - |
| GINA | - | 0.0% | - | - | - | - | - | - | - |
| HERB | - | 100.0% | - | - | - | - | - | - | - |
| ILBO | - | 0.0% | - | - | - | - | - | - | - |
| JANU | - | 100.0% | - | - | - | - | - | - | - |
| JOUS | - | 100.0% | - | - | - | - | - | - | - |
| KEGA | - | 100.0% | - | - | - | - | - | - | - |
| LACA | - | 0.0% | - | - | - | - | - | - | - |
| LAIG | - | 100.0% | - | - | - | - | - | - | - |
| LAVL | - | 100.0% | - | - | - | - | - | - | - |
| LBRD | - | 0.0% | - | - | - | - | - | - | - |
| LCST | - | 100.0% | - | - | - | - | - | - | - |
| LCVA | - | 100.0% | - | - | - | - | - | - | - |
| LESA | - | 100.0% | - | - | - | - | - | - | - |
| LEZI | - | 100.0% | - | - | - | - | - | - | - |
| LIZ2 | - | 100.0% | - | - | - | - | - | - | - |
| LLGV | - | 100.0% | - | - | - | - | - | - | - |
| LMDM | - | 0.0% | - | - | - | - | - | - | - |
| LOYT | - | 100.0% | - | - | - | - | - | - | - |
| LROC | - | 0.0% | - | - | - | - | - | - | - |
| LUMI | - | 0.0% | - | - | - | - | - | - | - |
| MAGR | - | 100.0% | - | - | - | - | - | - | - |
| MAKS | - | 0.0% | - | - | - | - | - | - | - |
| MATC | - | 100.0% | - | - | - | - | - | - | - |
| MAUP | - | 100.0% | - | - | - | - | - | - | - |
| MAZE | - | 100.0% | - | - | - | - | - | - | - |
| MGIS | - | 0.0% | - | - | - | - | - | - | - |
| MTP2 | - | 0.0% | - | - | - | - | - | - | - |
| NGER | - | 100.0% | - | - | - | - | - | - | - |
| NMTR | - | 100.0% | - | - | - | - | - | - | - |
| NOGT | - | 100.0% | - | - | - | - | - | - | - |
| NOYL | - | 100.0% | - | - | - | - | - | - | - |
| NVPT | - | 100.0% | - | - | - | - | - | - | - |
| OMBA | - | 100.0% | - | - | - | - | - | - | - |
| PALI | - | 0.0% | - | - | - | - | - | - | - |
| PAMO | - | 100.0% | - | - | - | - | - | - | - |
| PAYE | - | 0.0% | - | - | - | - | - | - | - |
| PEY2 | - | 100.0% | - | - | - | - | - | - | - |
| PLGN | - | 100.0% | - | - | - | - | - | - | - |
| PLMG | - | 0.0% | - | - | - | - | - | - | - |
| PLST | - | 100.0% | - | - | - | - | - | - | - |
| PNTV | - | 100.0% | - | - | - | - | - | - | - |
| POBU | - | 0.0% | - | - | - | - | - | - | - |
| PRR2 | - | 100.0% | - | - | - | - | - | - | - |
| PTRC | - | 100.0% | - | - | - | - | - | - | - |
| PUYA | - | 100.0% | - | - | - | - | - | - | - |
| ROSD | - | 100.0% | - | - | - | - | - | - | - |
| ROSI | - | 0.0% | - | - | - | - | - | - | - |
| SBL2 | - | 0.0% | - | - | - | - | - | - | - |
| SCOA | - | 0.0% | - | - | - | - | - | - | - |
| SCOP | - | 0.0% | - | - | - | - | - | - | - |
| SEM2 | - | 100.0% | - | - | - | - | - | - | - |
| SJDV | - | 0.0% | - | - | - | - | - | - | - |
| SMCR | - | 100.0% | - | - | - | - | - | - | - |
| SMLE | - | 0.0% | - | - | - | - | - | - | - |
| SPIO | - | 100.0% | - | - | - | - | - | - | - |
| STAZ | - | 100.0% | - | - | - | - | - | - | - |
| STF2 | - | 100.0% | - | - | - | - | - | - | - |
| STGN | - | 100.0% | - | - | - | - | - | - | - |
| STMR | - | 0.0% | - | - | - | - | - | - | - |
| VDOM | - | 0.0% | - | - | - | - | - | - | - |
| VNTE | - | 100.0% | - | - | - | - | - | - | - |
| VOUZ | - | 100.0% | - | - | - | - | - | - | - |
| VRCT | - | 100.0% | - | - | - | - | - | - | - |
| VSOU | - | 100.0% | - | - | - | - | - | - | - |
| WARE | - | 0.0% | - | - | - | - | - | - | - |
| ZIMM | - | 0.0% | - | - | - | - | - | - | - |
| GRAS | x | 80.0% | 80.6% | - | 5.9168m | - | 14 | 1 | x |
| MODA | x | 100.0% | 84% | - | 6.9108m | - | 0 | 0 | x |
| STJ9 | x | 100.0% | 84.9% | - | 7.0275m | - | 30 | 39 | x |
| VILD | x | 100.0% | 86.3% | - | 7.8046m | - | 5 | 6 | x |
| RMZT | x | 100.0% | 86.5% | - | 4.1311m | - | 0 | 0 | x |
| FIED | x | 100.0% | 86.6% | - | 6.3130m | - | 27 | 47 | x |
| DOJX | x | 100.0% | 87.9% | - | 8.3986m | - | 20 | 1 | x |
| FLRC | x | 100.0% | 88.1% | - | 6.6981m | - | 1 | 2 | x |
| PIAA | x | 100.0% | 88.6% | - | 6.7721m | - | 2 | 3 | x |
| MSGT | x | 100.0% | 89.4% | - | 7.2418m | - | 0 | 1 | x |
| ELBA | x | 100.0% | 89.8% | - | 7.3086m | - | 52 | 10 | x |
| OSAN | x | 100.0% | 90.3% | - | 6.2843m | - | 0 | 0 | x |
| CDBC | x | 100.0% | 90.7% | - | 7.9719m | - | 24 | 16 | x |
| FILF | x | 100.0% | 90.8% | - | 6.0996m | - | 9 | 6 | x |
| CRTE | x | 100.0% | 90.9% | - | 9.3765m | - | 6 | 3 | x |
| GENO | x | 100.0% | 90.9% | - | 7.7714m | - | 43 | 38 | x |
| MARG | x | 100.0% | 91% | - | 7.7055m | - | 2 | 0 | x |
| BOUS | x | 100.0% | 91.3% | - | 7.5555m | - | 2 | 3 | x |
| CROL | x | 100.0% | 91.5% | - | 7.5096m | - | 3 | 0 | x |
| MAUL | x | 100.0% | 91.9% | - | 7.0929m | - | 33 | 48 | x |
| RIXH | x | 100.0% | 92.1% | - | 8.6994m | - | 8 | 12 | x |
| LUCE | x | 100.0% | 92.4% | - | 12.4041m | - | 18 | 1 | x |
| CHTG | x | 100.0% | 92.6% | - | 9.8156m | - | 118 | 13 | x |
| LAHE | x | 100.0% | 92.6% | - | 6.5325m | - | 204 | 74 | x |
| STNZ | x | 100.0% | 93% | - | 6.0220m | - | 3 | 1 | x |
| TORI | x | 100.0% | 93.3% | - | 7.7320m | - | 28 | 32 | x |
| TUDA | x | 100.0% | 93.5% | - | 6.8923m | - | 0 | 1 | x |
| BRDO | x | 100.0% | 93.6% | - | 9.0832m | - | 4 | 0 | x |
| COVI | x | 100.0% | 93.6% | - | 7.6505m | - | 12 | 17 | x |
| MSMM | x | 100.0% | 93.7% | - | 7.7890m | - | 19 | 5 | x |
| CAMA | x | 100.0% | 94% | - | 6.5995m | - | 25 | 5 | x |
| RNNE | x | 100.0% | 94% | - | 7.4836m | - | 41 | 8 | x |
| SOLR | x | 100.0% | 94.2% | - | 7.6953m | - | 4 | 0 | x |
| BLG2 | x | 100.0% | 94.3% | - | 8.1233m | - | 10 | 1 | x |
| MELN | x | 100.0% | 94.3% | - | 8.6227m | - | 1 | 1 | x |
| BACT | x | 100.0% | 94.4% | - | 6.8549m | - | 1 | 0 | x |
| MRON | x | 100.0% | 94.6% | - | 6.4406m | - | 3 | 4 | x |
| BSCN | x | 100.0% | 94.7% | - | 7.9900m | - | 57 | 37 | x |
| MDEU | x | 100.0% | 94.7% | - | 16.4647m | - | 50 | 11 | x |
| SAFF | x | 100.0% | 94.7% | - | 7.0315m | - | 2 | 0 | x |
| SARZ | x | 100.0% | 94.7% | - | 8.0360m | - | 8 | 5 | x |
| BUSI | x | 100.0% | 94.8% | - | 6.7157m | - | 0 | 6 | x |
| NIVO | x | 100.0% | 94.9% | - | 8.3396m | - | 4 | 6 | x |
| ANGL | x | 100.0% | 95.1% | - | 7.3221m | - | 3 | 2 | x |
| VIRG | x | 100.0% | 95.1% | - | 10.3164m | - | 122 | 8 | x |
| DOCO | x | 100.0% | 95.2% | - | 8.2855m | - | 99 | 35 | x |
| PUYV | x | 100.0% | 95.2% | - | 6.5583m | - | 20 | 20 | x |
| BEAV | x | 100.0% | 95.4% | - | 12.2337m | - | 9 | 4 | x |
| AGDS | x | 100.0% | 95.5% | - | 6.4416m | - | 0 | 1 | x |
| ERCK | x | 100.0% | 95.5% | - | 10.2781m | - | 14 | 4 | x |
| FRTT | x | 100.0% | 95.5% | - | 8.4034m | - | 7 | 2 | x |
| MARS | x | 100.0% | 95.5% | - | 7.1053m | - | 46 | 2 | x |
| MPRV | x | 100.0% | 95.5% | - | 6.2468m | - | 0 | 1 | x |
| SGIL | x | 100.0% | 95.5% | - | 7.4658m | - | 124 | 30 | x |
| MICH | x | 100.0% | 95.6% | - | 7.3818m | - | 22 | 3 | x |
| THNV | x | 100.0% | 95.6% | - | 6.2445m | - | 0 | 0 | x |
| DIPP | x | 100.0% | 95.8% | - | 9.1682m | - | 85 | 18 | x |
| ROMY | x | 100.0% | 95.9% | - | 5.6688m | - | 0 | 1 | x |
| RUPT | x | 100.0% | 95.9% | - | 4.6560m | - | 0 | 0 | x |
| CEVY | x | 100.0% | 96.1% | - | 5.1070m | - | 30 | 2 | x |
| CHTA | x | 100.0% | 96.1% | - | 9.0108m | - | 23 | 5 | x |
| LPPZ | x | 100.0% | 96.1% | - | 7.4096m | - | 7 | 0 | x |
| MTMN | x | 100.0% | 96.1% | - | 7.6455m | - | 20 | 5 | x |
| ATH2 | x | 100.0% | 96.2% | - | 6.5633m | - | 0 | 8 | x |
| BAUG | x | 100.0% | 96.2% | - | 6.3130m | - | 0 | 0 | x |
| CANT | x | 100.0% | 96.2% | - | 9.5574m | - | 23 | 22 | x |
| CBRY | x | 100.0% | 96.2% | - | 8.2935m | - | 14 | 0 | x |
| EZEV | x | 100.0% | 96.2% | - | 8.2009m | - | 12 | 1 | x |
| AMB2 | x | 100.0% | 96.3% | - | 7.6121m | - | 7 | 2 | x |
| APT2 | x | 100.0% | 96.3% | - | 5.1225m | - | 0 | 0 | x |
| LURI | x | 100.0% | 96.3% | - | 7.4386m | - | 0 | 1 | x |
| NARB | x | 100.0% | 96.3% | - | 6.7075m | - | 52 | 11 | x |
| OISO | x | 100.0% | 96.3% | - | 8.2120m | - | 46 | 0 | x |
| SIST | x | 100.0% | 96.3% | - | 4.7494m | - | 0 | 2 | x |
| TRYS | x | 100.0% | 96.3% | - | 6.5987m | - | 0 | 7 | x |
| LUCI | x | 100.0% | 96.4% | - | 6.7892m | - | 0 | 0 | x |
| MLLC | x | 100.0% | 96.4% | - | 6.5854m | - | 2 | 1 | x |
| NICA | x | 100.0% | 96.4% | - | 7.4277m | - | 0 | 2 | x |
| STV2 | x | 100.0% | 96.4% | - | 8.0097m | - | 5 | 2 | x |
| ALU2 | x | 100.0% | 96.5% | - | 5.7739m | - | 0 | 0 | x |
| ATST | x | 100.0% | 96.5% | - | 7.6600m | - | 37 | 1 | x |
| BIAZ | x | 100.0% | 96.5% | - | 6.3452m | - | 6 | 1 | x |
| CHBR | x | 100.0% | 96.5% | - | 7.9902m | - | 24 | 3 | x |
| LLIV | x | 100.0% | 96.5% | - | 6.4259m | - | 20 | 12 | x |
| PRAC | x | 100.0% | 96.5% | - | 6.2408m | - | 35 | 7 | x |
| QNSN | x | 100.0% | 96.5% | - | 5.7731m | - | 0 | 0 | x |
| RAYL | x | 100.0% | 96.5% | - | 7.4735m | - | 15 | 0 | x |
| RST2 | x | 100.0% | 96.5% | - | 5.5814m | - | 68 | 3 | x |
| SAMR | x | 100.0% | 96.5% | - | 7.3225m | - | 0 | 1 | x |
| TRBL | x | 100.0% | 96.5% | - | 5.6911m | - | 0 | 10 | x |
| AXPV | x | 100.0% | 96.6% | - | 8.5568m | - | 56 | 0 | x |
| BRST | x | 100.0% | 96.6% | - | 9.4087m | - | 42 | 3 | x |
| CAMP | x | 100.0% | 96.6% | - | 6.9037m | - | 7 | 0 | x |
| CANE | x | 100.0% | 96.6% | - | 7.8618m | - | 0 | 0 | x |
| CUBX | x | 100.0% | 96.6% | - | 7.8883m | - | 41 | 9 | x |
| FAYE | x | 100.0% | 96.6% | - | 7.5272m | - | 23 | 3 | x |
| GRAC | x | 100.0% | 96.6% | - | 6.9126m | - | 58 | 0 | x |
| MNBL | x | 100.0% | 96.6% | - | 4.6306m | - | 0 | 0 | x |
| SAFQ | x | 100.0% | 96.6% | - | 6.0813m | - | 0 | 0 | x |
| SARL | x | 100.0% | 96.6% | - | 9.1475m | - | 27 | 1 | x |
| SARS | x | 100.0% | 96.6% | - | 6.6738m | - | 0 | 0 | x |
| SNNT | x | 100.0% | 96.6% | - | 6.2512m | - | 0 | 2 | x |
| TLTG | x | 100.0% | 96.6% | - | 10.6220m | - | 23 | 0 | x |
| VEN3 | x | 100.0% | 96.6% | - | 6.5651m | - | 0 | 0 | x |
| VISN | x | 100.0% | 96.6% | - | 7.8450m | - | 2 | 0 | x |
| ALGY | x | 100.0% | 96.7% | - | 4.8652m | - | 0 | 2 | x |
| ANDE | x | 100.0% | 96.7% | - | 5.7375m | - | 0 | 0 | x |
| ARDN | x | 100.0% | 96.7% | - | 5.5376m | - | 0 | 0 | x |
| ASNE | x | 100.0% | 96.7% | - | 4.4774m | - | 0 | 0 | x |
| BARD | x | 100.0% | 96.7% | - | 5.9292m | - | 0 | 0 | x |
| BRG9 | x | 100.0% | 96.7% | - | 4.5011m | - | 0 | 0 | x |
| BRM2 | x | 100.0% | 96.7% | - | 6.6872m | - | 0 | 2 | x |
| CACI | x | 100.0% | 96.7% | - | 8.0485m | - | 15 | 0 | x |
| CHDY | x | 100.0% | 96.7% | - | 8.3194m | - | 2 | 2 | x |
| CLFD | x | 100.0% | 96.7% | - | 5.7949m | - | 0 | 0 | x |
| DOUA | x | 100.0% | 96.7% | - | 8.6265m | - | 12 | 9 | x |
| ETOI | x | 100.0% | 96.7% | - | 10.3063m | - | 66 | 22 | x |
| ISLA | x | 100.0% | 96.7% | - | 8.1397m | - | 4 | 2 | x |
| MOFA | x | 100.0% | 96.7% | - | 6.8702m | - | 0 | 2 | x |
| MONS | x | 100.0% | 96.7% | - | 6.4260m | - | 0 | 0 | x |
| PZNA | x | 100.0% | 96.7% | - | 7.1668m | - | 43 | 0 | x |
| QUYC | x | 100.0% | 96.7% | - | 5.2361m | - | 0 | 0 | x |
| SCDA | x | 100.0% | 96.7% | - | 7.1821m | - | 10 | 0 | x |
| SET1 | x | 100.0% | 96.7% | - | 6.6438m | - | 70 | 0 | x |
| TGRI | x | 100.0% | 96.7% | - | 6.4809m | - | 0 | 0 | x |
| VERN | x | 100.0% | 96.7% | - | 5.3024m | - | 0 | 0 | x |
| VIN1 | x | 100.0% | 96.7% | - | 6.4795m | - | 0 | 0 | x |
| AIGL | x | 100.0% | 96.8% | - | 4.1445m | - | 87 | 0 | x |
| AMNS | x | 100.0% | 96.8% | - | 6.0500m | - | 0 | 0 | x |
| AUNI | x | 100.0% | 96.8% | - | 5.8129m | - | 0 | 0 | x |
| BAL2 | x | 100.0% | 96.8% | - | 7.0731m | - | 0 | 0 | x |
| BEUG | x | 100.0% | 96.8% | - | 6.8665m | - | 0 | 0 | x |
| BRIV | x | 100.0% | 96.8% | - | 6.4033m | - | 42 | 9 | x |
| C2CN | x | 100.0% | 96.8% | - | 6.2209m | - | 0 | 0 | x |
| CERN | x | 100.0% | 96.8% | - | 7.8658m | - | 15 | 2 | x |
| DOUL | x | 100.0% | 96.8% | - | 8.4534m | - | 2 | 0 | x |
| EGLT | x | 100.0% | 96.8% | - | 6.2322m | - | 40 | 0 | x |
| GDIJ | x | 100.0% | 96.8% | - | 5.9088m | - | 0 | 0 | x |
| JOI2 | x | 100.0% | 96.8% | - | 7.1844m | - | 13 | 1 | x |
| LGES | x | 100.0% | 96.8% | - | 6.4183m | - | 1 | 0 | x |
| MIRE | x | 100.0% | 96.8% | - | 6.1552m | - | 0 | 0 | x |
| MOMO | x | 100.0% | 96.8% | - | 6.4474m | - | 0 | 0 | x |
| MTIL | x | 100.0% | 96.8% | - | 4.5752m | - | 0 | 0 | x |
| MULH | x | 100.0% | 96.8% | - | 8.3751m | - | 11 | 1 | x |
| OPME | x | 100.0% | 96.8% | - | 6.9471m | - | 0 | 1 | x |
| RYO2 | x | 100.0% | 96.8% | - | 5.8983m | - | 0 | 0 | x |
| SAUN | x | 100.0% | 96.8% | - | 5.4703m | - | 0 | 0 | x |
| STBX | x | 100.0% | 96.8% | - | 6.2492m | - | 65 | 7 | x |
| STMA | x | 100.0% | 96.8% | - | 6.6013m | - | 0 | 0 | x |
| TANZ | x | 100.0% | 96.8% | - | 9.7166m | - | 15 | 3 | x |
| TRMO | x | 100.0% | 96.8% | - | 7.0673m | - | 16 | 2 | x |
| UZER | x | 100.0% | 96.8% | - | 5.9061m | - | 0 | 0 | x |
| VAUD | x | 100.0% | 96.8% | - | 6.2199m | - | 0 | 0 | x |
| VLPL | x | 100.0% | 96.8% | - | 5.2777m | - | 0 | 1 | x |
| ANAY | x | 100.0% | 96.9% | - | 9.2260m | - | 90 | 2 | x |
| ARCI | x | 100.0% | 96.9% | - | 7.9566m | - | 2 | 0 | x |
| AVAL | x | 100.0% | 96.9% | - | 7.0211m | - | 49 | 6 | x |
| AVRA | x | 100.0% | 96.9% | - | 7.5395m | - | 3 | 0 | x |
| BLVR | x | 100.0% | 96.9% | - | 8.1653m | - | 33 | 2 | x |
| BMHG | x | 100.0% | 96.9% | - | 8.5231m | - | 54 | 0 | x |
| BRMF | x | 100.0% | 96.9% | - | 7.1469m | - | 73 | 0 | x |
| BRMZ | x | 100.0% | 96.9% | - | 8.4033m | - | 7 | 0 | x |
| CHIZ | x | 100.0% | 96.9% | - | 9.1729m | - | 42 | 1 | x |
| COAU | x | 100.0% | 96.9% | - | 9.0172m | - | 3 | 1 | x |
| CRHX | x | 100.0% | 96.9% | - | 8.1685m | - | 14 | 1 | x |
| CRSM | x | 100.0% | 96.9% | - | 8.1863m | - | 31 | 0 | x |
| DGLG | x | 100.0% | 96.9% | - | 8.5247m | - | 94 | 9 | x |
| DUNQ | x | 100.0% | 96.9% | - | 10.6051m | - | 49 | 3 | x |
| FLER | x | 100.0% | 96.9% | - | 7.9551m | - | 14 | 1 | x |
| FOUG | x | 100.0% | 96.9% | - | 8.3672m | - | 40 | 3 | x |
| GLRA | x | 100.0% | 96.9% | - | 7.2691m | - | 6 | 0 | x |
| GORN | x | 100.0% | 96.9% | - | 7.8848m | - | 17 | 0 | x |
| LONS | x | 100.0% | 96.9% | - | 7.9194m | - | 23 | 10 | x |
| LPAS | x | 100.0% | 96.9% | - | 7.4437m | - | 13 | 0 | x |
| MAN2 | x | 100.0% | 96.9% | - | 7.2294m | - | 20 | 1 | x |
| OTER | x | 100.0% | 96.9% | - | 8.1654m | - | 6 | 0 | x |
| PAYR | x | 100.0% | 96.9% | - | 7.8415m | - | 3 | 0 | x |
| RGNC | x | 100.0% | 96.9% | - | 7.3394m | - | 0 | 3 | x |
| SEUR | x | 100.0% | 96.9% | - | 8.2644m | - | 29 | 1 | x |
| SOUS | x | 100.0% | 96.9% | - | 7.4108m | - | 3 | 1 | x |
| SRMG | x | 100.0% | 96.9% | - | 7.7072m | - | 8 | 3 | x |
| UCAG | x | 100.0% | 96.9% | - | 7.5935m | - | 40 | 0 | x |
| UNIX | x | 100.0% | 96.9% | - | 7.4111m | - | 6 | 0 | x |
| VOUR | x | 100.0% | 96.9% | - | 7.9422m | - | 81 | 0 | x |
| AGLM | x | 100.0% | 97% | - | 7.8514m | - | 18 | 1 | x |
| AICI | x | 100.0% | 97% | - | 4.6217m | - | 0 | 1 | x |
| AILT | x | 100.0% | 97% | - | 8.5260m | - | 15 | 0 | x |
| ANGE | x | 100.0% | 97% | - | 7.1297m | - | 2 | 0 | x |
| AUTN | x | 100.0% | 97% | - | 8.5976m | - | 15 | 0 | x |
| BOGU | x | 100.0% | 97% | - | 8.0548m | - | 11 | 1 | x |
| BREV | x | 100.0% | 97% | - | 7.8517m | - | 3 | 0 | x |
| CALS | x | 100.0% | 97% | - | 8.3210m | - | 51 | 0 | x |
| CASE | x | 100.0% | 97% | - | 7.2556m | - | 30 | 0 | x |
| CAUD | x | 100.0% | 97% | - | 9.0077m | - | 1 | 2 | x |
| CEPI | x | 100.0% | 97% | - | 6.1441m | - | 38 | 1 | x |
| CHAS | x | 100.0% | 97% | - | 8.6633m | - | 7 | 0 | x |
| CHBS | x | 100.0% | 97% | - | 7.9860m | - | 3 | 0 | x |
| CHIO | x | 100.0% | 97% | - | 6365348.2083m | - | 31 | 0 | x |
| CHLN | x | 100.0% | 97% | - | 8.1866m | - | 21 | 0 | x |
| CHRE | x | 100.0% | 97% | - | 8.4695m | - | 4 | 0 | x |
| CHRX | x | 100.0% | 97% | - | 8.2170m | - | 16 | 0 | x |
| CLFE | x | 100.0% | 97% | - | 6.1240m | - | 2 | 0 | x |
| CNTL | x | 100.0% | 97% | - | 8.6063m | - | 32 | 0 | x |
| CRVA | x | 100.0% | 97% | - | 7.2466m | - | 7 | 0 | x |
| DAGO | x | 100.0% | 97% | - | 8.3670m | - | 46 | 2 | x |
| DIPL | x | 100.0% | 97% | - | 7.1546m | - | 45 | 0 | x |
| DROU | x | 100.0% | 97% | - | 8.6710m | - | 29 | 0 | x |
| DRUS | x | 100.0% | 97% | - | 9.0221m | - | 5 | 0 | x |
| ENTZ | x | 100.0% | 97% | - | 8.6677m | - | 51 | 0 | x |
| EPR2 | x | 100.0% | 97% | - | 8.0404m | - | 3 | 0 | x |
| FCMP | x | 100.0% | 97% | - | 8.4648m | - | 8 | 0 | x |
| FDET | x | 100.0% | 97% | - | 6.9067m | - | 101 | 2 | x |
| FEGA | x | 100.0% | 97% | - | 8.0420m | - | 41 | 0 | x |
| FET2 | x | 100.0% | 97% | - | 8.2563m | - | 4 | 0 | x |
| FOU2 | x | 100.0% | 97% | - | 7.7492m | - | 9 | 0 | x |
| GRON | x | 100.0% | 97% | - | 8.1531m | - | 9 | 0 | x |
| GUIP | x | 100.0% | 97% | - | 8.7928m | - | 41 | 0 | x |
| HERS | x | 100.0% | 97% | - | 8.6190m | - | 14 | 0 | x |
| HOLA | x | 100.0% | 97% | - | 6.9908m | - | 24 | 0 | x |
| HONF | x | 100.0% | 97% | - | 8.4908m | - | 57 | 3 | x |
| HOUL | x | 100.0% | 97% | - | 8.4574m | - | 129 | 0 | x |
| HRSN | x | 100.0% | 97% | - | 7.7052m | - | 4 | 0 | x |
| IGNP | x | 100.0% | 97% | - | 10.4051m | - | 7 | 0 | x |
| ILDX | x | 100.0% | 97% | - | 7.8069m | - | 60 | 0 | x |
| IRAF | x | 100.0% | 97% | - | 6.1601m | - | 30 | 10 | x |
| JARG | x | 100.0% | 97% | - | 8.6165m | - | 27 | 0 | x |
| JONZ | x | 100.0% | 97% | - | 8.6688m | - | 25 | 0 | x |
| KARL | x | 100.0% | 97% | - | 9.2613m | - | 55 | 0 | x |
| KONE | x | 100.0% | 97% | - | 8.0465m | - | 22 | 0 | x |
| LBUG | x | 100.0% | 97% | - | 8.1584m | - | 7 | 0 | x |
| LENE | x | 100.0% | 97% | - | 8.1279m | - | 6 | 0 | x |
| LETO | x | 100.0% | 97% | - | 7.8782m | - | 45 | 1 | x |
| LGBO | x | 100.0% | 97% | - | 7.8356m | - | 7 | 0 | x |
| LIL2 | x | 100.0% | 97% | - | 8.0930m | - | 43 | 0 | x |
| LULI | x | 100.0% | 97% | - | 7.5502m | - | 116 | 5 | x |
| MACH | x | 100.0% | 97% | - | 7.2863m | - | 4 | 0 | x |
| MLVL | x | 100.0% | 97% | - | 7.7890m | - | 22 | 0 | x |
| MTBT | x | 100.0% | 97% | - | 8.5316m | - | 25 | 0 | x |
| NACY | x | 100.0% | 97% | - | 8.9291m | - | 5 | 0 | x |
| NAYC | x | 100.0% | 97% | - | 6.1985m | - | 87 | 2 | x |
| NMCU | x | 100.0% | 97% | - | 6.4490m | - | 2 | 0 | x |
| PERP | x | 100.0% | 97% | - | 8.2022m | - | 6 | 0 | x |
| PLEM | x | 100.0% | 97% | - | 7.5159m | - | 6 | 0 | x |
| PMTH | x | 100.0% | 97% | - | 6365662.9298m | - | 37 | 0 | x |
| PRNY | x | 100.0% | 97% | - | 7.7787m | - | 67 | 0 | x |
| REDO | x | 100.0% | 97% | - | 7.6867m | - | 7 | 0 | x |
| RENN | x | 100.0% | 97% | - | 8.0176m | - | 5 | 0 | x |
| ROTG | x | 100.0% | 97% | - | 11.0722m | - | 37 | 2 | x |
| ROYA | x | 100.0% | 97% | - | 7.9075m | - | 4 | 0 | x |
| SEES | x | 100.0% | 97% | - | 7.7627m | - | 15 | 0 | x |
| SIRT | x | 100.0% | 97% | - | 7.0744m | - | 0 | 0 | x |
| SJDA | x | 100.0% | 97% | - | 6.8499m | - | 49 | 0 | x |
| SJPL | x | 100.0% | 97% | - | 6.9336m | - | 79 | 0 | x |
| SMSP | x | 100.0% | 97% | - | 9.0940m | - | 5 | 0 | x |
| STBR | x | 100.0% | 97% | - | 7.6036m | - | 8 | 1 | x |
| STJF | x | 100.0% | 97% | - | 9.3329m | - | 49 | 0 | x |
| STLO | x | 100.0% | 97% | - | 7.8223m | - | 6 | 0 | x |
| STPS | x | 100.0% | 97% | - | 8.1604m | - | 1 | 0 | x |
| THIL | x | 100.0% | 97% | - | 7.9003m | - | 8 | 0 | x |
| VFCH | x | 100.0% | 97% | - | 7.2382m | - | 34 | 1 | x |
| YPOR | x | 100.0% | 97% | - | 8.0031m | - | 37 | 0 | x |
| ZARA | x | 100.0% | 97% | - | 9.2665m | - | 9 | 0 | x |
| AGN2 | x | 100.0% | 97.1% | - | 4.8781m | - | 0 | 0 | x |
| CAPT | x | 100.0% | 97.1% | - | 4.0920m | - | 0 | 0 | x |
| STGS | x | 100.0% | 97.1% | - | 3.9606m | - | 0 | 0 | x |
| STS2 | x | 100.0% | 97.1% | - | 3.8028m | - | 0 | 0 | x |
| LCAN | x | 100.0% | 97.2% | - | 9.3707m | - | 76 | 10 | x |
| TLSE | x | 100.0% | 97.2% | - | 7.5671m | - | 19 | 1 | x |
| AUCH | x | 100.0% | 97.3% | - | 7.2931m | - | 12 | 0 | x |
| BMNT | x | 100.0% | 97.3% | - | 6.5299m | - | 47 | 0 | x |
| CRAL | x | 100.0% | 97.3% | - | 6.0544m | - | 22 | 0 | x |
| ESCO | x | 100.0% | 97.3% | - | 5.6785m | - | 19 | 2 | x |
| EUSK | x | 100.0% | 97.3% | - | 10.0317m | - | 182 | 0 | x |
| FAJP | x | 100.0% | 97.3% | - | 6.7245m | - | 23 | 0 | x |
| LACO | x | 100.0% | 97.3% | - | 7.2917m | - | 14 | 0 | x |
| LGAR | x | 100.0% | 97.3% | - | 7.4381m | - | 29 | 0 | x |
| MTDM | x | 100.0% | 97.3% | - | 7.6588m | - | 10 | 0 | x |
| PESS | x | 100.0% | 97.3% | - | 7.6328m | - | 10 | 0 | x |
| PUY2 | x | 100.0% | 97.3% | - | 6.8371m | - | 1 | 0 | x |
| RIO1 | x | 100.0% | 97.3% | - | 6.4071m | - | 15 | 0 | x |
| TLMF | x | 100.0% | 97.3% | - | 6.3639m | - | 49 | 0 | x |
| TLSG | x | 100.0% | 97.3% | - | 7.2055m | - | 5 | 0 | x |
| UNME | x | 100.0% | 97.3% | - | 6.4240m | - | 11 | 1 | x |




Calcul des coordonnées sur le code
Les stations sont éliminées du calcul si l'écart entre les coordonnées issues du calcul sur le code et les coordonnées initiales est supérieur à dE=10.0 m, dN=10.0 m ou dU=20.0 m| Station | dE (m) | dN (m) | dU (m) |
|---|---|---|---|
| AGDS | -0.35m | 0.10m | 0.52m |
| AGLM | -0.45m | 0.32m | 0.35m |
| AGN2 | -0.18m | -0.03m | 0.57m |
| AICI | -0.13m | 0.21m | 0.43m |
| AIGL | -0.20m | 0.30m | 0.76m |
| AILT | -0.44m | 0.18m | 0.54m |
| ALGY | -0.47m | 0.19m | 0.36m |
| ALU2 | -0.14m | 0.12m | 0.64m |
| AMB2 | -0.48m | 0.36m | 0.18m |
| AMNS | -0.27m | 0.06m | 0.29m |
| ANAY | -0.56m | 0.16m | 0.57m |
| ANDE | -0.16m | -0.03m | 0.46m |
| ANGE | -0.40m | 0.12m | 0.87m |
| ANGL | -0.35m | 0.37m | 0.85m |
| APT2 | -0.13m | 0.30m | -0.54m |
| ARCI | -0.38m | 0.27m | 0.76m |
| ARDN | -0.20m | -0.02m | 0.12m |
| ASNE | -0.13m | 0.12m | 0.53m |
| ATH2 | -0.38m | 0.09m | 0.87m |
| ATST | -0.39m | 0.41m | -0.14m |
| AUCH | -0.27m | 0.21m | 0.97m |
| AUNI | -0.28m | 0.01m | 0.65m |
| AUTN | -0.59m | 0.17m | 0.36m |
| AVAL | -0.48m | 0.15m | 0.76m |
| AVRA | -0.38m | 0.30m | 0.85m |
| AXPV | -0.63m | 0.37m | -0.11m |
| BACT | -0.41m | 0.37m | -0.13m |
| BAL2 | -0.44m | 0.08m | 0.57m |
| BARD | -0.25m | 0.41m | 0.23m |
| BAUG | -0.19m | 0.11m | 0.44m |
| BEAV | -0.45m | 0.25m | 0.77m |
| BEUG | -0.38m | 0.08m | 0.64m |
| BIAZ | -0.34m | 0.41m | 0.65m |
| BLG2 | -0.57m | 0.48m | 0.33m |
| BLVR | -0.32m | 0.25m | 0.59m |
| BMHG | -0.38m | 0.28m | 0.44m |
| BMNT | -0.41m | 0.13m | 0.82m |
| BOGU | -0.46m | 0.15m | 0.32m |
| BOUS | -0.37m | 0.26m | 0.70m |
| BRDO | -0.39m | 0.15m | 0.03m |
| BREV | -0.34m | 0.27m | 0.91m |
| BRG9 | -0.44m | 0.31m | 0.33m |
| BRIV | -0.64m | 0.19m | 1.15m |
| BRM2 | -0.39m | 0.18m | 0.45m |
| BRMF | -0.61m | 0.16m | 0.36m |
| BRMZ | -0.48m | 0.33m | 0.74m |
| BRST | -0.32m | 0.07m | 0.82m |
| BSCN | -0.47m | 0.35m | 0.35m |
| BUSI | -0.29m | -0.02m | 0.55m |
| C2CN | -0.40m | 0.30m | 0.55m |
| CACI | -0.56m | 0.21m | 0.27m |
| CALS | -0.39m | 0.15m | 0.88m |
| CAMA | -0.48m | 0.26m | 0.10m |
| CAMP | -0.35m | 0.05m | 0.41m |
| CANE | -0.45m | 0.22m | 0.45m |
| CANT | -0.42m | 0.31m | 0.84m |
| CAPT | -0.34m | 0.05m | 0.41m |
| CASE | -0.42m | 0.25m | 0.61m |
| CAUD | -0.45m | 0.18m | 0.53m |
| CBRY | -0.36m | 0.38m | 0.32m |
| CDBC | -0.35m | 0.40m | 0.84m |
| CEPI | -0.55m | 0.24m | 0.68m |
| CERN | -0.37m | 0.03m | 0.51m |
| CEVY | -0.45m | 0.22m | 0.63m |
| CHAS | -0.38m | 0.10m | 0.64m |
| CHBR | -0.34m | 0.13m | 0.65m |
| CHBS | -0.28m | -0.15m | 0.62m |
| CHDY | -0.45m | 0.18m | 0.56m |
| CHIO | -0.38m | 0.12m | 1.18m |
| CHIZ | -0.43m | 0.31m | 0.90m |
| CHLN | -0.27m | 0.07m | 0.22m |
| CHRE | -0.35m | 0.13m | 0.72m |
| CHRX | -0.38m | 0.23m | 0.89m |
| CHTA | -0.17m | 0.32m | 0.83m |
| CHTG | -0.41m | 0.31m | 1.85m |
| CLFD | -0.48m | 0.25m | 0.89m |
| CLFE | -0.39m | 0.09m | 0.50m |
| CNTL | -0.35m | 0.15m | 1.19m |
| COAU | -0.48m | 0.22m | 0.47m |
| COVI | -0.32m | 0.15m | 1.04m |
| CRAL | -0.36m | 0.33m | 0.88m |
| CRHX | -0.38m | 0.52m | 0.59m |
| CROL | -0.44m | 0.01m | 0.44m |
| CRSM | -0.44m | 0.10m | 0.95m |
| CRTE | -0.57m | 0.07m | -0.11m |
| CRVA | -0.38m | 0.15m | 0.73m |
| CUBX | -0.47m | 0.35m | 1.14m |
| DAGO | -0.47m | 0.09m | 0.31m |
| DGLG | -0.36m | 0.12m | 0.55m |
| DIPL | -0.22m | 0.18m | 0.90m |
| DIPP | -0.40m | 0.14m | 0.87m |
| DOCO | -0.41m | 0.09m | 0.64m |
| DOJX | -0.52m | 0.33m | 0.27m |
| DOUA | -0.36m | 0.08m | 0.66m |
| DOUL | -0.43m | 0.19m | 0.78m |
| DROU | -0.22m | 0.05m | 0.51m |
| DRUS | -0.44m | 0.13m | 0.47m |
| DUNQ | -0.41m | 0.07m | 0.75m |
| EGLT | -0.44m | 0.19m | 0.36m |
| ELBA | -0.50m | 0.15m | -0.12m |
| ENTZ | -0.46m | 0.09m | 0.31m |
| EPR2 | -0.45m | 0.16m | 0.89m |
| ERCK | -0.64m | 0.08m | 0.93m |
| ESCO | -0.41m | 0.26m | 0.57m |
| ETOI | -0.36m | 0.15m | 0.48m |
| EUSK | -0.05m | -0.09m | 0.79m |
| EZEV | -0.42m | 0.37m | -0.14m |
| FAJP | -0.41m | 0.41m | 0.68m |
| FAYE | -0.42m | 0.33m | 0.02m |
| FCMP | -0.38m | 0.14m | 1.00m |
| FDET | -0.51m | 0.13m | 0.93m |
| FEGA | -0.43m | 0.22m | 0.63m |
| FET2 | -0.42m | 0.35m | 0.79m |
| FIED | -0.49m | 0.54m | 1.29m |
| FILF | -0.35m | 0.32m | 1.02m |
| FLER | -0.36m | 0.29m | 0.38m |
| FLRC | -0.68m | 0.34m | -0.12m |
| FOU2 | -0.20m | 0.08m | 0.83m |
| FOUG | -0.34m | 0.15m | 0.20m |
| FRTT | -0.45m | 0.21m | 0.50m |
| GDIJ | -0.12m | 0.19m | 0.10m |
| GENO | -0.25m | 0.31m | -0.07m |
| GLRA | -0.54m | 0.22m | 0.71m |
| GORN | -0.51m | 0.42m | 0.56m |
| GRAC | -0.69m | 0.35m | 0.40m |
| GRAS | -0.46m | 0.15m | 0.87m |
| GRON | -0.32m | 0.18m | 0.56m |
| GUIP | -0.60m | 0.42m | 0.83m |
| HERS | -0.07m | 0.02m | 0.73m |
| HOLA | -0.49m | 0.21m | 0.75m |
| HONF | -0.48m | 0.12m | 0.86m |
| HOUL | -0.40m | 0.11m | 0.84m |
| HRSN | -0.36m | 0.15m | 0.42m |
| IGNP | -0.05m | 0.03m | 0.31m |
| ILDX | -0.50m | 0.17m | 0.66m |
| IRAF | -0.49m | 0.27m | 0.74m |
| ISLA | -0.37m | 0.20m | 1.12m |
| JARG | -0.35m | 0.17m | 0.29m |
| JOI2 | -0.50m | 0.34m | 0.30m |
| JONZ | -0.34m | 0.08m | 0.80m |
| KARL | -0.01m | -0.10m | 0.06m |
| KONE | -0.38m | 0.34m | 0.44m |
| LACO | -0.42m | 0.33m | 0.66m |
| LAHE | -0.76m | -0.03m | 1.13m |
| LBUG | -0.53m | 0.16m | 0.79m |
| LCAN | -0.23m | 0.25m | 0.59m |
| LENE | -0.43m | 0.22m | 0.42m |
| LETO | -0.37m | -0.07m | 0.91m |
| LGAR | -0.40m | 0.33m | 0.09m |
| LGBO | -0.43m | 0.23m | 0.66m |
| LGES | -0.17m | 0.14m | 0.77m |
| LIL2 | -0.49m | 0.09m | 0.38m |
| LLIV | -0.46m | 0.35m | 0.50m |
| LONS | -0.46m | 0.31m | 0.29m |
| LPAS | -0.42m | 0.22m | 0.34m |
| LPPZ | -0.28m | 0.05m | 0.92m |
| LUCE | -0.72m | 0.38m | 1.81m |
| LUCI | -0.17m | 0.16m | -0.59m |
| LULI | -0.58m | 0.21m | 0.53m |
| LURI | -0.20m | 0.08m | 0.56m |
| MACH | -0.31m | 0.05m | 0.87m |
| MAN2 | -0.49m | 0.21m | 0.95m |
| MARG | -0.42m | 0.36m | -0.01m |
| MARS | -0.36m | 0.24m | -0.26m |
| MAUL | -0.20m | 0.33m | 0.14m |
| MDEU | -0.46m | 0.28m | -0.07m |
| MELN | -0.41m | 0.23m | 0.87m |
| MICH | -0.59m | -0.12m | 1.24m |
| MIRE | -0.29m | -0.03m | 0.54m |
| MLLC | -0.39m | 0.05m | 0.63m |
| MLVL | -0.49m | 0.13m | 0.50m |
| MNBL | -0.21m | 0.12m | -0.15m |
| MODA | -0.19m | 0.31m | 0.29m |
| MOFA | -0.34m | 0.04m | 0.81m |
| MOMO | -0.26m | 0.10m | 0.88m |
| MONS | -0.73m | 0.45m | 1.02m |
| MPRV | -0.35m | 0.23m | 0.68m |
| MRON | -0.54m | 0.12m | 0.46m |
| MSGT | -0.23m | 0.39m | 1.02m |
| MSMM | -0.37m | 0.49m | 0.14m |
| MTBT | -0.52m | 0.25m | 0.19m |
| MTDM | -0.34m | 0.30m | 1.11m |
| MTIL | -0.12m | 0.05m | 0.32m |
| MTMN | -0.17m | 0.45m | 0.24m |
| MULH | -0.37m | 0.12m | 0.53m |
| NACY | -0.40m | 0.08m | 0.68m |
| NARB | -0.39m | 0.29m | 0.27m |
| NAYC | -0.63m | 0.24m | 0.60m |
| NICA | -0.34m | 0.50m | -0.21m |
| NIVO | -0.54m | 0.26m | 0.68m |
| NMCU | 0.25m | -0.51m | -0.03m |
| OISO | -0.77m | 0.42m | 0.25m |
| OPME | -0.31m | 0.38m | 0.68m |
| OSAN | -0.49m | 0.28m | 0.11m |
| OTER | -0.42m | 0.57m | 0.10m |
| PAYR | -1.25m | 0.80m | 0.93m |
| PERP | -0.32m | 0.28m | 0.42m |
| PESS | -0.36m | 0.21m | 0.81m |
| PIAA | -0.40m | 0.33m | -0.27m |
| PLEM | -0.54m | 0.31m | 1.03m |
| PMTH | -0.47m | -0.04m | 1.28m |
| PRAC | -0.42m | 0.28m | 0.80m |
| PRNY | -0.40m | 0.16m | 0.73m |
| PUY2 | -0.28m | 0.41m | -0.07m |
| PUYV | -0.52m | 0.24m | 0.19m |
| PZNA | -0.19m | 0.25m | 0.43m |
| QNSN | -0.50m | 0.15m | 0.39m |
| QUYC | -0.25m | 0.04m | 0.36m |
| RAYL | -0.32m | 0.31m | -0.07m |
| REDO | -0.35m | 0.17m | 0.97m |
| RENN | -0.34m | 0.04m | 1.06m |
| RGNC | -0.30m | 0.20m | 0.51m |
| RIO1 | -0.45m | 0.26m | 0.68m |
| RIXH | -0.40m | 0.28m | 0.74m |
| RMZT | -0.39m | 0.11m | 0.23m |
| RNNE | -0.60m | 0.19m | 1.08m |
| ROMY | -0.35m | 0.07m | 0.39m |
| ROTG | -0.00m | 0.04m | 0.38m |
| ROYA | -0.32m | 0.19m | 0.74m |
| RST2 | -0.42m | 0.31m | 0.28m |
| RUPT | -0.34m | -0.03m | 0.59m |
| RYO2 | -0.42m | 0.09m | 0.80m |
| SAFF | -0.29m | 0.23m | 0.48m |
| SAFQ | -0.27m | 0.04m | 0.75m |
| SAMR | -0.34m | -0.01m | 0.81m |
| SARL | -0.45m | 0.23m | 0.32m |
| SARS | -0.41m | 0.16m | 0.04m |
| SARZ | -0.21m | 0.10m | 0.83m |
| SAUN | -0.33m | -0.16m | 0.12m |
| SCDA | -0.01m | -0.03m | 0.40m |
| SEES | -0.38m | 0.33m | 0.42m |
| SET1 | -0.43m | 0.13m | 0.20m |
| SEUR | -0.48m | 0.03m | 0.06m |
| SGIL | -0.45m | 0.28m | 0.20m |
| SIRT | -0.53m | 0.26m | 0.75m |
| SIST | -0.17m | 0.15m | -0.11m |
| SJDA | -0.35m | 0.22m | 0.93m |
| SJPL | -0.33m | 0.27m | 0.73m |
| SMSP | -0.39m | 0.12m | 0.63m |
| SNNT | -0.33m | 0.07m | 0.68m |
| SOLR | -0.33m | 0.36m | -0.42m |
| SOUS | -0.44m | 0.35m | 0.78m |
| SRMG | -0.30m | 0.40m | 0.34m |
| STBR | -0.58m | 0.23m | 0.91m |
| STBX | -0.41m | 0.51m | 0.41m |
| STGS | -0.26m | 0.01m | 0.15m |
| STJ9 | -0.39m | 0.66m | 0.59m |
| STJF | -0.41m | 0.24m | 0.89m |
| STLO | -0.35m | 0.27m | 0.68m |
| STMA | -0.40m | 0.32m | 0.50m |
| STNZ | 0.44m | -0.45m | 0.20m |
| STPS | -0.47m | 0.17m | 0.62m |
| STS2 | -0.34m | 0.26m | 0.13m |
| STV2 | -0.47m | 0.33m | 0.03m |
| TANZ | -0.46m | 0.33m | 0.43m |
| TGRI | -0.71m | 0.51m | 1.17m |
| THIL | -0.35m | 0.17m | 0.59m |
| THNV | -0.33m | -0.04m | 0.30m |
| TLMF | -0.58m | 0.16m | 0.67m |
| TLSE | -0.47m | 0.00m | 1.51m |
| TLSG | -0.17m | -0.02m | 0.83m |
| TLTG | -0.05m | 0.05m | -0.47m |
| TORI | -0.36m | 0.28m | 0.28m |
| TRBL | -0.38m | 0.16m | 0.73m |
| TRMO | -0.44m | 0.37m | 0.33m |
| TRYS | -0.38m | 0.10m | 0.64m |
| TUDA | -0.31m | 0.28m | 0.19m |
| UCAG | -0.29m | 0.32m | -0.11m |
| UNIX | -0.50m | 0.19m | 0.11m |
| UNME | -0.25m | 0.39m | 1.03m |
| UZER | -0.15m | 0.20m | 0.01m |
| VAUD | -0.52m | 0.12m | 0.39m |
| VEN3 | -0.16m | 0.09m | 1.22m |
| VERN | -0.25m | 0.18m | 0.56m |
| VFCH | -0.55m | 0.26m | 0.43m |
| VILD | -0.51m | 0.18m | 0.39m |
| VIN1 | -0.68m | 0.52m | 1.30m |
| VIRG | -0.35m | 0.35m | -0.12m |
| VISN | -0.33m | 0.30m | 0.24m |
| VLPL | -0.06m | 0.12m | 0.75m |
| VOUR | -0.23m | 0.15m | 0.63m |
| YPOR | -0.54m | -0.05m | 0.86m |
| ZARA | -0.48m | 0.30m | 0.85m |




Fixation des ambiguités
| Station 1 | Station 2 | Longueur | Constellation | Ambiguités posées | Ambiguités fixées QIF | Ambiguités fixées L12 | Ratio |
|---|---|---|---|---|---|---|---|
| GRAC | GRAS | 0.032km | G | 40 | 28 | 12 | 100.000% |
| OSAN | PIAA | 9.910km | G | 32 | 26 | 4 | 93.750% |
| APT2 | RST2 | 10.538km | G | 38 | 28 | 10 | 100.000% |
| BRDO | LURI | 12.642km | G | 40 | 30 | 8 | 95.000% |
| MSMM | QNSN | 16.958km | G | 44 | 28 | 14 | 95.455% |
| MICH | RST2 | 18.862km | G | 46 | 28 | 13 | 89.130% |
| ATST | CRTE | 20.958km | G | 34 | 26 | - | 76.5% |
| BRDO | TUDA | 21.050km | G | 36 | 28 | - | 77.8% |
| EZEV | NICA | 23.146km | G | 46 | 30 | - | 65.2% |
| CANE | GRAC | 23.531km | G | 44 | 32 | - | 72.7% |
| CANE | NICA | 23.758km | G | 44 | 26 | - | 59.1% |
| FAYE | GRAC | 23.864km | G | 42 | 30 | - | 71.4% |
| LUCI | TUDA | 27.929km | G | 40 | 28 | - | 70.0% |
| ATST | LUCI | 29.073km | G | 44 | 28 | - | 63.6% |
| CAMA | RAYL | 29.272km | G | 36 | 28 | - | 77.8% |
| MICH | QNSN | 33.591km | G | 44 | 28 | - | 63.6% |
| ATST | SARS | 34.652km | G | 42 | 32 | - | 76.2% |
| MICH | SIST | 37.330km | G | 72 | 30 | - | 41.7% |
| CAMP | PIAA | 37.642km | G | 42 | 26 | - | 61.9% |
| CAMA | FAYE | 38.152km | G | 38 | 30 | - | 78.9% |
| SIST | STV2 | 40.585km | G | 58 | 30 | - | 51.7% |
| CRTE | OSAN | 43.003km | G | 34 | 32 | - | 94.1% |
| APT2 | AXPV | 43.016km | G | 36 | 28 | - | 77.8% |
| CAMA | QNSN | 43.800km | G | 34 | 30 | - | 88.2% |
| CAMA | TLTG | 46.042km | G | 38 | 28 | - | 73.7% |
| BACT | STV2 | 47.604km | G | 36 | 30 | - | 83.3% |
| CAMP | MAUL | 48.537km | G | 42 | 26 | - | 61.9% |
| BRDO | ELBA | 60.313km | G | 34 | 28 | - | 82.4% |
| MAUL | MDEU | 70.520km | G | 40 | 28 | - | 70.0% |
| ELBA | VIRG | 100.349km | G | 36 | 26 | - | 72.2% |
| EZEV | GENO | 134.664km | G | 40 | 30 | - | 75.0% |
| GENO | VIRG | 154.028km | G | 40 | 26 | - | 65.0% |
| MDEU | UCAG | 184.570km | G | 40 | 30 | - | 75.0% |
| AXPV | MARS | 23.663km | G | 36 | 26 | - | 72.2% |
| CLFD | CLFE | 3.623km | G | 38 | 32 | 6 | 100.000% |
| CLFD | OPME | 5.600km | G | 46 | 30 | 16 | 100.000% |
| BRMF | VOUR | 14.218km | G | 48 | 30 | 14 | 91.667% |
| FIED | LONS | 15.984km | G | 50 | 26 | 23 | 98.000% |
| MRON | PRNY | 17.571km | G | 48 | 32 | 12 | 91.667% |
| FIED | VAUD | 24.129km | G | 50 | 24 | - | 48.0% |
| AUTN | CACI | 29.413km | G | 42 | 30 | - | 71.4% |
| OPME | SJPL | 31.660km | G | 44 | 32 | - | 72.7% |
| RMZT | SOLR | 32.558km | G | 34 | 28 | - | 82.4% |
| CBRY | CROL | 33.780km | G | 38 | 26 | - | 68.4% |
| BRMF | NIVO | 34.630km | G | 42 | 32 | - | 76.2% |
| RMZT | VISN | 34.746km | G | 36 | 28 | - | 77.8% |
| CROL | VILD | 35.288km | G | 40 | 32 | - | 80.0% |
| DROU | SEUR | 35.428km | G | 42 | 34 | - | 81.0% |
| SEUR | VAUD | 36.191km | G | 42 | 34 | - | 81.0% |
| ANAY | UNIX | 37.412km | G | 50 | 36 | - | 72.0% |
| GDIJ | SEUR | 38.490km | G | 48 | 30 | - | 62.5% |
| BLG2 | CERN | 38.716km | G | 44 | 32 | - | 72.7% |
| CERN | MARG | 39.704km | G | 44 | 32 | - | 72.7% |
| SOLR | VILD | 41.742km | G | 34 | 26 | - | 76.5% |
| AMB2 | UNIX | 42.944km | G | 38 | 34 | - | 89.5% |
| ANAY | GLRA | 46.276km | G | 40 | 32 | - | 80.0% |
| CBRY | NIVO | 46.852km | G | 44 | 30 | - | 68.2% |
| ANAY | VOUR | 47.459km | G | 38 | 32 | - | 84.2% |
| AUTN | DROU | 47.925km | G | 50 | 30 | - | 60.0% |
| FIED | MRON | 48.802km | G | 52 | 28 | - | 53.8% |
| PUYV | UNIX | 49.694km | G | 42 | 32 | - | 76.2% |
| AMB2 | CLFE | 53.362km | G | 34 | 32 | - | 94.1% |
| PUYV | VERN | 54.386km | G | 46 | 34 | - | 73.9% |
| BLG2 | LONS | 56.342km | G | 40 | 30 | - | 75.0% |
| SOLR | STV2 | 58.442km | G | 38 | 32 | - | 84.2% |
| AMB2 | RNNE | 61.026km | G | 30 | 28 | - | 93.3% |
| CROL | MODA | 64.868km | G | 32 | 22 | - | 68.8% |
| BLG2 | BRMF | 69.887km | G | 40 | 30 | - | 75.0% |
| MODA | TORI | 76.675km | G | 40 | 28 | - | 70.0% |
| TLSE | TLSG | 1.266km | G | 52 | 28 | 24 | 100.000% |
| SAFF | SAFQ | 2.717km | G | 40 | 30 | 10 | 100.000% |
| AGDS | PZNA | 15.801km | G | 40 | 26 | 10 | 90.000% |
| AGDS | SET1 | 20.829km | G | 50 | 30 | - | 60.0% |
| CEPI | FAJP | 21.558km | G | 42 | 34 | - | 81.0% |
| AIGL | FLRC | 22.705km | G | 50 | 30 | - | 60.0% |
| AGN2 | LGAR | 23.213km | G | 48 | 30 | - | 62.5% |
| HOLA | SAFF | 26.190km | G | 42 | 32 | - | 76.2% |
| CRAL | STGS | 26.785km | G | 48 | 30 | - | 62.5% |
| AIGL | HOLA | 33.408km | G | 44 | 32 | - | 72.7% |
| FILF | LLIV | 37.619km | G | 46 | 26 | - | 56.5% |
| SAFQ | TRMO | 37.889km | G | 44 | 32 | - | 72.7% |
| LACO | STGS | 38.675km | G | 46 | 38 | - | 82.6% |
| FILF | PERP | 40.759km | G | 42 | 28 | - | 66.7% |
| AUCH | BMNT | 40.760km | G | 56 | 36 | - | 64.3% |
| AGDS | NARB | 42.678km | G | 48 | 28 | - | 58.3% |
| LACO | TLSG | 43.906km | G | 38 | 32 | - | 84.2% |
| AGN2 | BMNT | 43.972km | G | 50 | 32 | - | 64.0% |
| FAJP | PAYR | 45.918km | G | 38 | 34 | - | 89.5% |
| MTIL | PRAC | 48.200km | G | 52 | 32 | - | 61.5% |
| FLRC | STBX | 48.638km | G | 44 | 30 | - | 68.2% |
| FAJP | MSGT | 48.679km | G | 38 | 36 | - | 94.7% |
| ESCO | STGS | 51.626km | G | 48 | 32 | - | 66.7% |
| LLIV | MSGT | 52.764km | G | 38 | 30 | - | 78.9% |
| BMNT | TLSE | 53.716km | G | 52 | 32 | - | 61.5% |
| BMNT | MTIL | 55.833km | G | 68 | 36 | - | 52.9% |
| NARB | PERP | 57.039km | G | 40 | 34 | - | 85.0% |
| ESCO | MSGT | 57.393km | G | 40 | 32 | - | 80.0% |
| FLRC | SCDA | 58.268km | G | 40 | 28 | - | 70.0% |
| HOLA | PZNA | 60.030km | G | 46 | 26 | - | 56.5% |
| SET1 | SGIL | 66.696km | G | 48 | 26 | - | 54.2% |
| PRAC | SOUS | 77.929km | G | 38 | 34 | - | 89.5% |
| CASE | FILF | 85.324km | G | 38 | 32 | - | 84.2% |
| ESCO | ZARA | 193.529km | G | 50 | 30 | - | 60.0% |
| TLMF | TLSE | 8.686km | G | 50 | 36 | 13 | 98.000% |
| CHBS | VFCH | 6.813km | G | 36 | 32 | 4 | 100.000% |
| ARDN | JARG | 20.796km | G | 38 | 28 | - | 73.7% |
| SIRT | IGNP | 21.675km | G | 42 | 30 | - | 71.4% |
| BOUS | SNNT | 23.295km | G | 46 | 28 | - | 60.9% |
| ALU2 | CACI | 23.317km | G | 42 | 34 | - | 81.0% |
| BRMZ | MAN2 | 24.118km | G | 40 | 34 | - | 85.0% |
| MONS | STPS | 26.700km | G | 44 | 30 | - | 68.2% |
| AILT | VIN1 | 27.842km | G | 44 | 34 | - | 77.3% |
| FDET | RGNC | 30.122km | G | 58 | 30 | - | 51.7% |
| FDET | SAUN | 30.296km | G | 44 | 32 | - | 72.7% |
| AVAL | VIN1 | 32.008km | G | 38 | 30 | - | 78.9% |
| LULI | MONS | 32.467km | G | 44 | 30 | - | 68.2% |
| JARG | OISO | 32.628km | G | 44 | 28 | - | 63.6% |
| OTER | RGNC | 35.116km | G | 50 | 30 | - | 60.0% |
| MELN | IGNP | 38.500km | G | 38 | 26 | - | 68.4% |
| BRG9 | LGES | 43.739km | G | 44 | 36 | - | 81.8% |
| ARCI | TRBL | 45.321km | G | 46 | 40 | - | 87.0% |
| ARCI | BRMZ | 45.582km | G | 38 | 30 | - | 78.9% |
| GRON | LULI | 46.301km | G | 44 | 32 | - | 72.7% |
| ASNE | JARG | 47.327km | G | 42 | 30 | - | 71.4% |
| MTMN | NAYC | 47.493km | G | 44 | 28 | - | 63.6% |
| CHBS | STMA | 47.614km | G | 38 | 30 | - | 78.9% |
| ASNE | GRON | 47.709km | G | 42 | 32 | - | 76.2% |
| OTER | STMA | 48.202km | G | 46 | 30 | - | 65.2% |
| AVAL | CACI | 48.613km | G | 40 | 32 | - | 80.0% |
| BOUS | BRG9 | 52.009km | G | 38 | 28 | - | 73.7% |
| NAYC | OTER | 54.539km | G | 44 | 30 | - | 68.2% |
| ASNE | VFCH | 54.793km | G | 50 | 34 | - | 68.0% |
| BAUG | FDET | 55.484km | G | 46 | 34 | - | 73.9% |
| FEGA | MELN | 56.020km | G | 38 | 28 | - | 73.7% |
| BAUG | MAN2 | 56.625km | G | 58 | 36 | - | 62.1% |
| ALU2 | MONS | 60.719km | G | 38 | 30 | - | 78.9% |
| BRG9 | EGLT | 66.011km | G | 40 | 30 | - | 75.0% |
| SIRT | TRBL | 66.027km | G | 62 | 34 | - | 54.8% |
| SNNT | STPS | 71.125km | G | 50 | 32 | - | 64.0% |
| REDO | SAMR | 5.304km | G | 52 | 30 | 21 | 98.077% |
| BRST | GUIP | 9.498km | G | 48 | 38 | 10 | 100.000% |
| LAHE | HONF | 14.986km | G | 46 | 30 | 15 | 97.826% |
| CRSM | EPR2 | 15.171km | G | 36 | 32 | 3 | 97.222% |
| AVRA | VLPL | 18.696km | G | 58 | 30 | 28 | 100.000% |
| BRST | LPPZ | 20.853km | G | 42 | 34 | - | 81.0% |
| EPR2 | HOUL | 24.167km | G | 36 | 32 | - | 88.9% |
| HOUL | LAHE | 24.992km | G | 46 | 34 | - | 73.9% |
| MACH | NMCU | 30.954km | G | 40 | 38 | - | 95.0% |
| STLO | VLPL | 34.876km | G | 62 | 30 | - | 48.4% |
| LPAS | SARZ | 36.676km | G | 52 | 32 | - | 61.5% |
| PLEM | STBR | 39.079km | G | 36 | 28 | - | 77.8% |
| FLER | GORN | 41.694km | G | 44 | 32 | - | 72.7% |
| REDO | STNZ | 42.414km | G | 48 | 44 | - | 91.7% |
| CHBR | COVI | 43.233km | G | 46 | 32 | - | 69.6% |
| MACH | STNZ | 44.497km | G | 48 | 44 | - | 91.7% |
| GUIP | ROTG | 44.840km | G | 56 | 34 | - | 60.7% |
| CHBR | RENN | 48.522km | G | 36 | 32 | - | 88.9% |
| CRSM | STLO | 49.617km | G | 34 | 26 | - | 76.5% |
| CHBR | SAMR | 50.123km | G | 56 | 36 | - | 64.3% |
| AVRA | GORN | 50.759km | G | 44 | 34 | - | 77.3% |
| DIPL | STBR | 50.859km | G | 42 | 32 | - | 76.2% |
| JOI2 | MACH | 51.073km | G | 34 | 30 | - | 88.2% |
| SARZ | STNZ | 51.249km | G | 58 | 36 | - | 62.1% |
| CRHX | KONE | 52.269km | G | 34 | 30 | - | 88.2% |
| AVRA | DIPL | 53.532km | G | 42 | 30 | - | 71.4% |
| COVI | GORN | 53.639km | G | 46 | 30 | - | 65.2% |
| FLER | SEES | 55.780km | G | 46 | 32 | - | 69.6% |
| CRHX | ROTG | 57.270km | G | 44 | 34 | - | 77.3% |
| ANGE | COVI | 58.381km | G | 48 | 32 | - | 66.7% |
| CRHX | STBR | 65.263km | G | 46 | 32 | - | 69.6% |
| BMHG | STLO | 82.434km | G | 48 | 28 | - | 58.3% |
| BMHG | CHIO | 168.038km | G | 40 | 30 | - | 75.0% |
| BMHG | PMTH | 184.821km | G | 46 | 34 | - | 73.9% |
| BMHG | CHTG | 14.047km | G | 64 | 36 | 24 | 93.750% |
| FCMP | YPOR | 4.899km | G | 38 | 34 | 4 | 100.000% |
| DGLG | DUNQ | 6.246km | G | 56 | 36 | 17 | 94.643% |
| BOGU | CNTL | 6.737km | G | 44 | 34 | 9 | 97.727% |
| LENE | VEN3 | 10.385km | G | 50 | 30 | 17 | 94.000% |
| BUSI | CAUD | 11.213km | G | 54 | 30 | 21 | 94.444% |
| IGNP | MLVL | 11.918km | G | 38 | 32 | 6 | 100.000% |
| ATH2 | COAU | 13.352km | G | 44 | 32 | 10 | 95.455% |
| CDBC | STJF | 15.516km | G | 36 | 28 | 5 | 91.667% |
| CDBC | MOFA | 16.842km | G | 40 | 26 | 12 | 95.000% |
| FCMP | MOFA | 18.192km | G | 46 | 34 | 11 | 97.826% |
| FET2 | ROMY | 19.149km | G | 48 | 32 | 11 | 89.583% |
| HONF | STJF | 21.154km | G | 36 | 32 | - | 88.9% |
| DOUA | LIL2 | 24.378km | G | 42 | 36 | - | 85.7% |
| CDBC | CNTL | 24.616km | G | 40 | 32 | - | 80.0% |
| FOU2 | MOMO | 28.158km | G | 36 | 32 | - | 88.9% |
| BOGU | MOMO | 29.212km | G | 46 | 32 | - | 69.6% |
| CNTL | VEN3 | 30.356km | G | 50 | 30 | - | 60.0% |
| IGNP | ISLA | 32.986km | G | 42 | 32 | - | 76.2% |
| BREV | TGRI | 33.056km | G | 44 | 30 | - | 68.2% |
| BEUG | DOUA | 33.457km | G | 40 | 30 | - | 75.0% |
| AMNS | DOUL | 34.704km | G | 40 | 36 | - | 90.0% |
| ISLA | TGRI | 34.753km | G | 44 | 32 | - | 72.7% |
| CALS | DGLG | 35.346km | G | 46 | 36 | - | 78.3% |
| BEUG | CHLN | 36.076km | G | 42 | 34 | - | 81.0% |
| CAUD | DOUA | 37.082km | G | 46 | 30 | - | 65.2% |
| BEUG | DOUL | 37.680km | G | 40 | 34 | - | 85.0% |
| BEAV | ISLA | 38.808km | G | 38 | 30 | - | 78.9% |
| BREV | VEN3 | 43.380km | G | 52 | 32 | - | 61.5% |
| BUSI | COAU | 43.552km | G | 44 | 32 | - | 72.7% |
| ATH2 | FET2 | 43.849km | G | 46 | 30 | - | 65.2% |
| CRVA | FET2 | 43.946km | G | 44 | 32 | - | 72.7% |
| BUSI | HRSN | 45.012km | G | 48 | 34 | - | 70.8% |
| AMNS | BEAV | 46.939km | G | 46 | 30 | - | 65.2% |
| CALS | LETO | 52.605km | G | 38 | 34 | - | 89.5% |
| DOUL | LETO | 65.226km | G | 36 | 32 | - | 88.9% |
| HERS | LETO | 98.805km | G | 38 | 32 | - | 84.2% |
| DIPP | FOU2 | 36.962km | G | 52 | 30 | - | 57.7% |
| MULH | RIXH | 4.343km | G | 48 | 28 | 19 | 97.917% |
| DOJX | RUPT | 7.482km | G | 90 | 48 | 42 | 100.000% |
| ENTZ | STJ9 | 8.669km | G | 52 | 28 | 19 | 90.385% |
| BRM2 | STJ9 | 10.101km | G | 60 | 28 | 27 | 91.667% |
| CHRE | MIRE | 14.628km | G | 38 | 30 | 6 | 94.737% |
| THIL | ROMY | 16.340km | G | 44 | 38 | 4 | 95.455% |
| BRM2 | DRUS | 19.007km | G | 60 | 28 | 26 | 90.000% |
| BLVR | MNBL | 24.159km | G | 58 | 32 | - | 55.2% |
| ALGY | FRTT | 24.554km | G | 64 | 28 | - | 43.8% |
| BAL2 | RIXH | 25.113km | G | 42 | 28 | - | 66.7% |
| ERCK | SARL | 28.326km | G | 40 | 32 | - | 80.0% |
| DOCO | THNV | 30.225km | G | 48 | 28 | - | 58.3% |
| BRM2 | ERCK | 30.931km | G | 58 | 28 | - | 48.3% |
| ANDE | DAGO | 32.200km | G | 42 | 32 | - | 76.2% |
| LUCE | MULH | 33.713km | G | 38 | 28 | - | 73.7% |
| ENTZ | TANZ | 35.277km | G | 42 | 32 | - | 76.2% |
| LUCE | MNBL | 35.421km | G | 58 | 26 | - | 44.8% |
| BAL2 | TANZ | 36.488km | G | 46 | 32 | - | 69.6% |
| ALGY | BSCN | 37.489km | G | 60 | 32 | - | 53.3% |
| CHRE | NACY | 38.102km | G | 38 | 34 | - | 89.5% |
| ANDE | RUPT | 42.514km | G | 56 | 32 | - | 57.1% |
| MTBT | RUPT | 43.031km | G | 62 | 34 | - | 54.8% |
| DRUS | KARL | 43.317km | G | 48 | 40 | - | 83.3% |
| ANDE | SMSP | 45.251km | G | 36 | 32 | - | 88.9% |
| MTBT | TRYS | 46.345km | G | 68 | 34 | - | 50.0% |
| BLVR | BSCN | 47.816km | G | 56 | 30 | - | 53.6% |
| SMSP | THIL | 48.319km | G | 40 | 32 | - | 80.0% |
| FOUG | MIRE | 49.137km | G | 42 | 32 | - | 76.2% |
| CEVY | DOCO | 49.942km | G | 46 | 32 | - | 69.6% |
| DOCO | NACY | 51.086km | G | 54 | 28 | - | 51.9% |
| FOUG | MNBL | 54.413km | G | 46 | 30 | - | 65.2% |
| CEVY | SMSP | 54.898km | G | 38 | 30 | - | 78.9% |
| CEVY | CHDY | 60.540km | G | 42 | 30 | - | 71.4% |
| CHAS | TRYS | 62.991km | G | 56 | 32 | - | 57.1% |
| EUSK | THNV | 151.811km | G | 42 | 32 | - | 76.2% |
| ETOI | STJ9 | 7.383km | G | 52 | 26 | 22 | 92.308% |
| CUBX | PESS | 7.665km | G | 36 | 32 | 3 | 97.222% |
| CHRX | MPRV | 10.035km | G | 50 | 30 | 17 | 94.000% |
| MTDM | STS2 | 13.562km | G | 46 | 28 | 15 | 93.478% |
| AICI | MLLC | 16.545km | G | 46 | 32 | 10 | 91.304% |
| C2CN | SRMG | 19.565km | G | 42 | 34 | 5 | 92.857% |
| CHIZ | SJDA | 21.974km | G | 44 | 32 | - | 72.7% |
| AICI | PUY2 | 22.996km | G | 52 | 32 | - | 61.5% |
| IRAF | MLLC | 24.856km | G | 40 | 32 | - | 80.0% |
| BRIV | UZER | 29.404km | G | 60 | 40 | - | 66.7% |
| CHTA | RYO2 | 29.431km | G | 62 | 30 | - | 48.4% |
| QUYC | ROYA | 31.769km | G | 40 | 32 | - | 80.0% |
| ANGL | RYO2 | 32.018km | G | 62 | 30 | - | 48.4% |
| C2CN | LGAR | 33.888km | G | 38 | 32 | - | 84.2% |
| BARD | JONZ | 37.522km | G | 42 | 32 | - | 76.2% |
| UZER | EGLT | 38.124km | G | 38 | 26 | - | 68.4% |
| AUNI | SJDA | 38.131km | G | 46 | 30 | - | 65.2% |
| PUY2 | STS2 | 39.326km | G | 56 | 28 | - | 50.0% |
| AGLM | BARD | 39.994km | G | 44 | 32 | - | 72.7% |
| LCAN | PESS | 41.684km | G | 36 | 32 | - | 88.9% |
| LCAN | QUYC | 42.446km | G | 42 | 36 | - | 85.7% |
| JONZ | QUYC | 43.014km | G | 46 | 36 | - | 78.3% |
| AICI | BIAZ | 45.033km | G | 54 | 34 | - | 63.0% |
| AGLM | LGBO | 45.459km | G | 36 | 32 | - | 88.9% |
| MLLC | UNME | 46.054km | G | 42 | 28 | - | 66.7% |
| LBUG | SRMG | 47.802km | G | 36 | 34 | - | 94.4% |
| ANGL | AUNI | 48.668km | G | 38 | 28 | - | 73.7% |
| CAPT | MTDM | 49.341km | G | 40 | 30 | - | 75.0% |
| BRIV | LBUG | 50.475km | G | 42 | 34 | - | 81.0% |
| CAPT | LGAR | 50.721km | G | 38 | 32 | - | 84.2% |
| AUNI | ROYA | 52.051km | G | 42 | 34 | - | 81.0% |
| LBUG | LGBO | 55.647km | G | 36 | 34 | - | 94.4% |
| AGLM | CHRX | 57.570km | G | 36 | 30 | - | 83.3% |
| IRAF | RIO1 | 130.144km | G | 38 | 34 | - | 89.5% |
| CANT | RIO1 | 158.306km | G | 70 | 46 | - | 65.7% |
| AUNI | ILDX | 20.545km | G | 40 | 30 | - | 75.0% |




Fixation des ambiguités Core
| Station 1 | Station 2 | Longueur | Constellation | Ambiguités posées | Ambiguités fixées QIF | Ambiguités fixées L12 | Ratio |
|---|---|---|---|---|---|---|---|
| GRAS | NICA | 25.389km | G | 44 | 24 | - | 54.5% |
| MLVL | TGRI | 46.866km | G | 40 | 30 | - | 75.0% |
| TGRI | TRBL | 54.291km | G | 48 | 36 | - | 75.0% |
| DGLG | LIL2 | 69.994km | G | 54 | 36 | - | 66.7% |
| EGLT | LGES | 78.902km | G | 44 | 30 | - | 68.2% |
| ANGL | CHIZ | 82.692km | G | 38 | 30 | - | 78.9% |
| CLFD | EGLT | 91.721km | G | 40 | 28 | - | 70.0% |
| MLVL | ROMY | 94.021km | G | 46 | 30 | - | 65.2% |
| ROMY | TRYS | 99.431km | G | 60 | 28 | - | 46.7% |
| CLFD | PUYV | 99.856km | G | 42 | 28 | - | 66.7% |
| CRAL | TLSE | 102.285km | G | 44 | 30 | - | 68.2% |
| AIGL | PUYV | 105.192km | G | 54 | 28 | - | 51.9% |
| MAN2 | TRBL | 106.913km | G | 52 | 34 | - | 65.4% |
| BRMF | PUYV | 112.426km | G | 44 | 34 | - | 77.3% |
| ANGL | STNZ | 113.782km | G | 52 | 42 | - | 80.8% |
| AXPV | GRAS | 131.440km | G | 38 | 24 | - | 63.2% |
| CHIZ | LGES | 131.735km | G | 44 | 32 | - | 72.7% |
| AUTN | BSCN | 133.134km | G | 54 | 30 | - | 55.6% |
| ENTZ | THNV | 140.669km | G | 44 | 32 | - | 72.7% |
| CHIZ | CUBX | 141.095km | G | 40 | 30 | - | 75.0% |
| MAN2 | VFCH | 142.465km | G | 48 | 30 | - | 62.5% |
| AUTN | BRMF | 145.348km | G | 52 | 38 | - | 73.1% |
| AUTN | TRYS | 150.482km | G | 68 | 32 | - | 47.1% |
| GENO | NICA | 157.322km | G | 42 | 28 | - | 66.7% |
| AIGL | AXPV | 157.424km | G | 42 | 30 | - | 71.4% |
| LIL2 | ROMY | 166.466km | G | 48 | 30 | - | 62.5% |
| ROMY | THNV | 176.050km | G | 52 | 36 | - | 69.2% |
| AIGL | TLSE | 180.053km | G | 44 | 32 | - | 72.7% |
| BRST | STNZ | 211.442km | G | 52 | 46 | - | 88.5% |
| NICA | SARS | 270.967km | G | 46 | 30 | - | 65.2% |




Estimation des ZTD finaux (coordonnées fixées) - ADDNEQ2
| Statistiques | |
|---|---|
| Paramètres explicites | 8767 |
| Paramètres implicites | 1737 |
| Paramètres ajustés | 10504 |
| Observations | 269306 |
| Degré de liberté | 258802 |
| Chi**2/DOF | 1.73 |
| Stations | 0 |
| Fichier | Début | Fin | Obs. | Par. | Dof |
|---|---|---|---|---|---|
| MET_26136R_1.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 27664 | 2098 | 25566 |
| MET_26136R_2.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 28687 | 2137 | 26550 |
| MET_26136R_3.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 29387 | 2198 | 27189 |
| MET_26136R_4.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 30131 | 2193 | 27938 |
| MET_26136R_5.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 29405 | 2152 | 27253 |
| MET_26136R_6.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 31836 | 2238 | 29598 |
| MET_26136R_7.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 30115 | 2284 | 27831 |
| MET_26136R_8.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 30424 | 2167 | 28257 |
| MET_26136R_9.NQ0 | 2026-05-16 12:00:00 | 2026-05-17 00:00:00 | 24869 | 1925 | 22944 |